Paper: Billion-Scale Graph Foundation Models arXiv: 2602.04768 | February 2026 Authors: GraphBFF research team
Graph neural networks have a scaling problem that's different from the one facing language models. With language, you pre-train a transformer on text and fine-tune on downstream tasks — the architecture is universal, and the pre-training data (text on the internet) is abundant, relatively homogeneous, and easily processed. With graphs, every domain has a different schema: citation networks have paper nodes and author nodes and venue nodes with different feature spaces; molecular graphs have atom nodes and bond edges with chemical properties; social networks have user nodes and interaction edges with temporal features. Transferring across these domains has been theoretically attractive and practically miserable.
GraphBFF, introduced in February 2026, is the first paper to present a complete, reproducible recipe for building billion-parameter graph foundation models that work across arbitrary heterogeneous graphs. This is not just an incremental scaling result — it's a methodological contribution that addresses the fundamental obstacle to graph pre-training at scale.
The Problem: Heterogeneity at Every Level
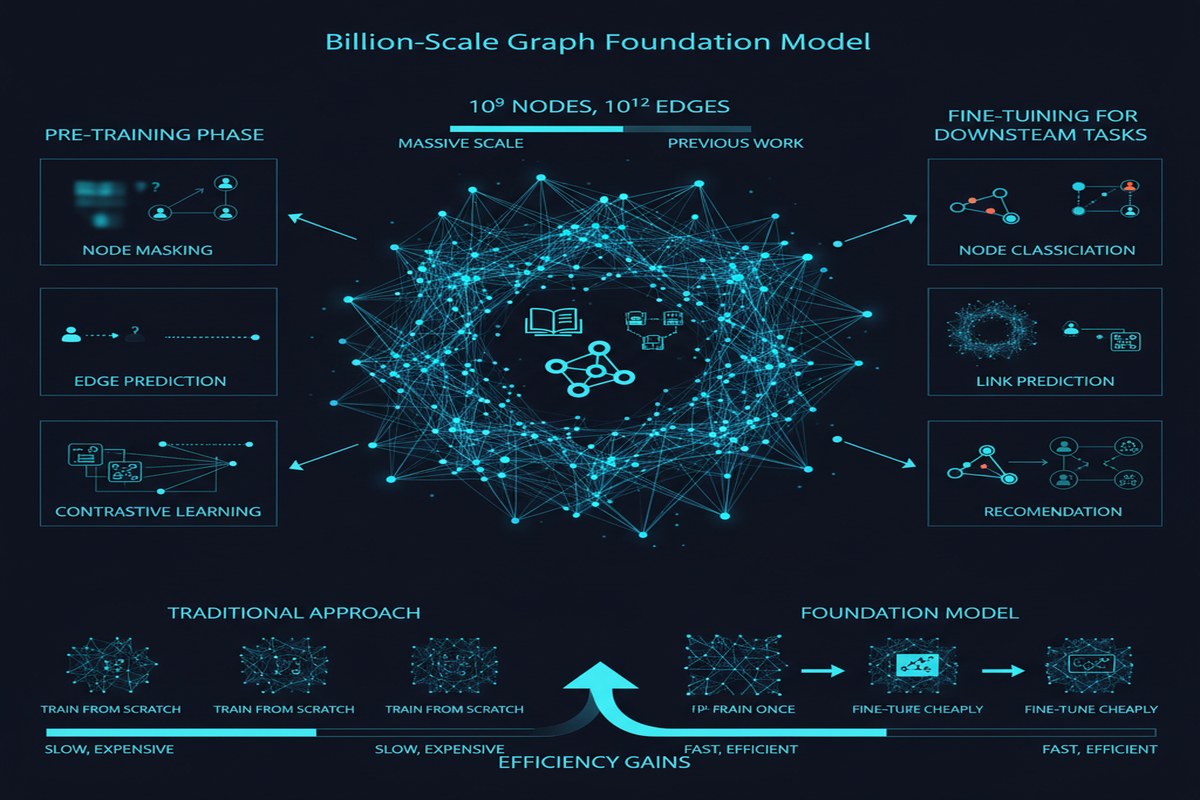
To appreciate why GraphBFF is hard, consider what "arbitrary heterogeneous graphs" actually means.
A typical knowledge graph (like Wikidata) has dozens of node types — Person, Organization, Location, Event, Concept — each with different feature schemas and different semantic roles. A social network has user nodes (with demographic features), content nodes (with text and media features), and interaction edges (with temporal and type information). A protein interaction network has protein nodes (with sequence and structure features) and interaction edges (with confidence scores and interaction type labels).
Pre-training a single model that can learn useful representations across all of these simultaneously requires solving several problems that have historically been treated separately:
- Feature space alignment: How do you handle nodes with different feature dimensionalities and types across graphs?
- Schema generalization: How do you encode the graph schema (what node types exist, what edge types connect them) in a way that transfers?
- Scale: How do you handle billion-node graphs that can't be loaded into memory simultaneously?
- Transfer learning: Pre-trained representations on one graph should be useful for downstream tasks on different graphs with different schemas.
The GraphBFF Architecture
GraphBFF addresses these challenges through a combination of architectural choices and pre-training strategies.
Tokenization for heterogeneous graphs: GraphBFF introduces a graph tokenization scheme that maps arbitrary node and edge features into a shared token space, using type-conditional encoders that learn to project diverse feature schemas into a common representation space. Think of it as the equivalent of a wordpiece tokenizer for graphs — a way to create a vocabulary that spans multiple domains.
The GraphBFF Transformer: The core architecture is a graph transformer with modifications for heterogeneous graphs. Rather than computing attention uniformly across all neighboring nodes, GraphBFF uses type-aware attention that distinguishes between different node and edge types. The model learns which types of neighbors are most informative for which types of queries — a citation paper attends differently to author neighbors than to venue neighbors.
Hierarchical sampling for billion-scale graphs: Billion-node graphs can't be processed with dense attention across all nodes simultaneously. GraphBFF uses a hierarchical neighborhood sampling strategy that preserves the statistical properties of the full graph while keeping the effective batch size tractable. The sampling strategy is designed to maintain long-range structural information that standard mini-batch GNN training loses.
graph TD
subgraph Input
G1[Heterogeneous Graph A\nCitation network]
G2[Heterogeneous Graph B\nSocial network]
G3[Heterogeneous Graph C\nProtein interactions]
end
subgraph Tokenization
T[Type-Conditional Encoders\nProject diverse features to shared space]
end
subgraph GraphBFF Transformer
SA[Type-Aware Self-Attention]
FFN[Feed-Forward Network]
HS[Hierarchical Sampling\nfor billion-scale]
end
subgraph Pre-training Objectives
MLM[Masked Node Prediction]
LPP[Link Presence Prediction]
GSR[Graph Structure Reconstruction]
end
subgraph Downstream Tasks
NC[Node Classification]
LP[Link Prediction]
GC[Graph Classification]
end
G1 --> T
G2 --> T
G3 --> T
T --> SA
SA --> FFN
HS --> SA
FFN --> MLM
FFN --> LPP
FFN --> GSR
FFN --> NC
FFN --> LP
FFN --> GC
Pre-training objectives: GraphBFF uses three complementary pre-training objectives: masked node prediction (reconstruct masked node features from neighbors), link presence prediction (predict whether an edge exists between two nodes), and graph structure reconstruction (recover structural properties from perturbed graphs). The combination ensures the model learns both feature-level and structure-level representations.
The Results
The empirical results demonstrate that billion-parameter graph models trained with GraphBFF's recipe outperform state-of-the-art GNN baselines across a diverse set of benchmark tasks, including node classification, link prediction, and graph-level classification across multiple domain-heterogeneous benchmarks.
More importantly, GraphBFF demonstrates genuine transfer learning: a model pre-trained on large heterogeneous graphs shows strong zero-shot and few-shot performance on held-out graphs with different schemas. This is the first convincing demonstration of this capability at billion-parameter scale.
The scaling behavior is also notable. Performance continues to improve as model parameters scale from 100M to 1B, following power-law scaling curves similar to what has been observed in language models. This suggests the approach is on a training trajectory where more compute will continue to yield improvements.
Why This Matters
Knowledge graphs are everywhere and underserved by current AI. Enterprises maintain enormous knowledge graphs — product catalogs, customer relationship graphs, supply chain networks, organizational hierarchies. The inability to build foundation models for these heterogeneous graph structures has been a genuine bottleneck. GraphBFF provides a viable path to the "pre-train once on your full graph, fine-tune for each downstream task" paradigm that has proven so powerful in NLP.
Biological networks are a high-value application. Drug discovery, protein interaction prediction, gene regulatory network analysis — all of these involve massive heterogeneous graphs where the node types are biochemically meaningful and the edges encode specific biological relationships. Foundation models for biological graphs that generalize across organisms, experimental conditions, and research questions could accelerate the kind of AI-assisted scientific discovery discussed in other posts.
The scaling law observation is significant. If billion-parameter graph models follow the same scaling laws as language models, then the trajectory is clear: larger models, more training data, better performance. The graph domain was stuck for years because it wasn't clear that scaling would work — this paper provides early evidence that it does.
Enterprise knowledge management. Every large organization has graph-structured knowledge that is currently underutilized because GNN tools require schema-specific training. A graph foundation model that can handle arbitrary schemas without retraining changes the economics of knowledge graph deployment entirely.
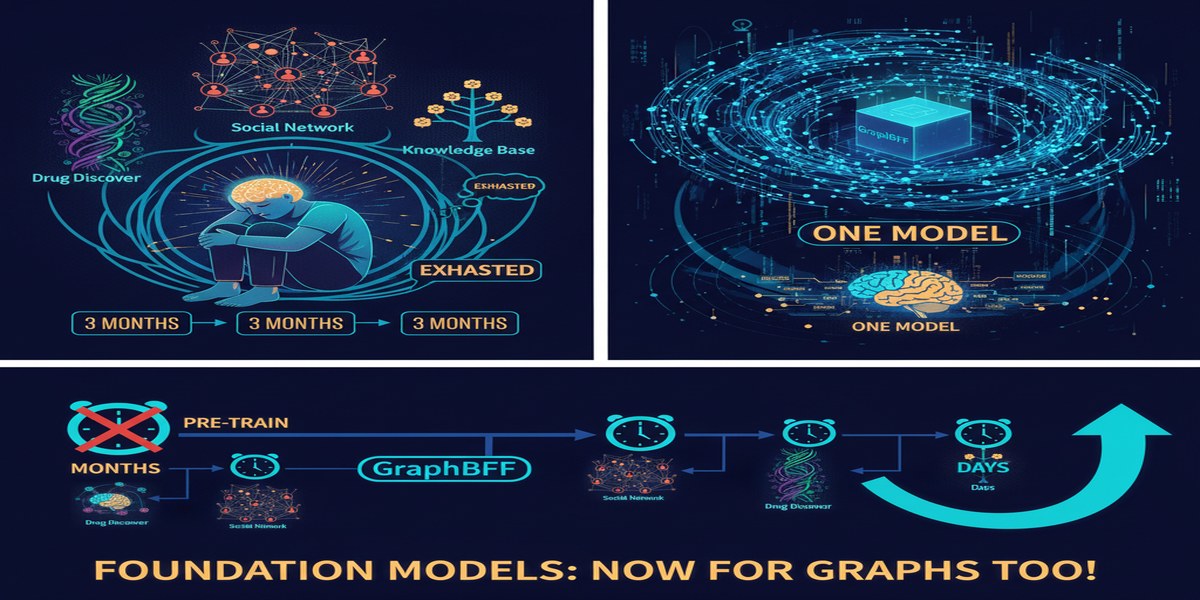
My Take
Graph neural networks have been "almost ready" for enterprise deployment for the better part of five years. The heterogeneity problem has been the primary obstacle — not the quality of GNN architectures, which are excellent for same-schema tasks, but the inability to transfer across domains that makes pre-training impractical.
GraphBFF doesn't fully solve this problem, but it provides the clearest path forward I've seen. The type-conditional tokenization and type-aware attention are exactly the right architectural primitives for handling heterogeneous schemas — analogous to how token type embeddings and segment encodings in BERT handled the heterogeneity between sentence-level context.
What I'd push back on: the scaling result is promising but not yet at the scale of the real production graphs I'm aware of. Billion-node graphs in the paper are research-scale; the largest enterprise knowledge graphs I've encountered have tens of billions of nodes and hundreds of billions of edges. The hierarchical sampling approach may not scale cleanly to that regime, and I'd want to see ablations at the production scale before betting heavily on this architecture.
The biological application is where I'm watching most closely. The combination of heterogeneous graph representations at scale with foundation model transfer learning is a capability set that doesn't exist today in the biological AI toolchain. If GraphBFF's approach generalizes to multi-omic networks — combining genomic, transcriptomic, proteomic, and metabolomic layers — the impact on drug discovery could be substantial.
The paper is technically dense but worth working through if you operate in any domain where graph-structured data is central to your problem. We've been waiting for this architecture for a long time.
arXiv:2602.04768 — read the full paper at arxiv.org/abs/2602.04768
Explore more from Dr. Jyothi



